Extracellular vesicles (EVs) are nano-sized lipid bilayer vesicles carrying various proteins, nucleic acids and small molecules. Secreted by all known cells, EVs are present in biofluids, making them readily accessible for study. EVs are internalised by cells, suggesting roles in cell-to-cell communication in both health and disease. Therefore, the uptake of EVs has become a major area of research, and an essential part of any study investigating EV functions. Due to the critical importance of EV uptake for function, tracking this phenomenon is one of the most common experiments presented in the EV literature. However, this cannot be done without first labelling EVs to enable visualisation and quantification.
What kinds of labels are used for visualising and tracking extracellular vesicles?
In vitro, EV uptake is usually measured using microscopy or flow cytometry, making fluorescence the label type of choice. For in vivo studies, fluorescence still tends to win out, but bioluminescence and radioactivity are used at times. Bioluminescence is commonly employed by transfecting the EV donor cell with the sequence for an EV marker conjugated to a bioluminescence-capable protein.1 The benefit of bioluminescence is that it is less prone to photobleaching than fluorescent labels, and so is more stable. However, due to the requirement for genetic modification of donor cells, bioluminescence labelling can only label EVs containing specific proteins. As EVs have no ubiquitous markers, the use of bioluminescence results in selective labelling. Radiolabelling EVs is rare, likely due to safety concerns and the resulting increased complexity of animal ethics applications.2 Given all of these considerations, the old faithful fluorescence remains the most popular choice for labelling EVs. But is this choice justified, and what are the challenges of common fluorescent labels?
A focus on fluorescent labels for extracellular vesicles
Fluorescent labels of any kind are hindered by the autofluorescence of cells, tissues and solutions, which can create a high level of background noise. Additionally, the propensity of fluorescent markers to photobleach and become unusable is problematic for many applications. This is exacerbated when the label strength is weak, as a longer exposure time is required – further increasing the risk of photobleaching.
Finally, weak tissue penetration makes fluorescence unsuitable or weakly sensitive for in vivo applications, often resulting in the need for ex vivo organ dissection for localisation of uptake. Despite these fairly major drawbacks, fluorescence is still the method of choice for labelling EVs, likely due to the familiarity of fluorescence and the availability of instruments for detection.
Lipophilic dyes
The most commonly used EV labels are the lipophilic fluorescent dyes. These primarily include the Sigma-Aldrich (i.e., Merck)-owned PKH family (named for their discoverer Paul Karl Horan, if you were wondering) and Invitrogen’s series of long chain carbocyanines, such as DiI (somewhat comically pronounced ‘Dye Aye’, Newcastle style). Despite differences in chemical structure, all lipophilic dyes work similarly to each other, intercalating into lipid bilayers such as those of EVs by virtue of their hydrophobic tails. The protocol for labelling EVs with lipophilic dyes is simple. The required concentration dye is mixed with your EV isolate and incubated for the desired length of time at the desired temperature. Often, though not always, excess dye is sequestered by the addition of EV-depleted fetal bovine serum (FBS) or bovine serum albumin (BSA).
Non-lipophilic dyes
Non-lipophilic fluorescent dyes used less commonly for EV labelling include CellMask Orange (which labels F-actin)3 and CFSE (which labels intraluminal amines)4. The uniformity of labelling with these dyes is less well studied than lipophilic dyes, perhaps due to their lower popularity. However, it could reasonably be assumed that different EVs (e.g., different subtypes or cells of origin) may have different F-actin or intraluminal amine levels, which would make these labelling methods more intrinsically variable and biased than lipophilic markers which would be expected to label all EVs equally well (or equally poorly…).
Labelling extracellular markers
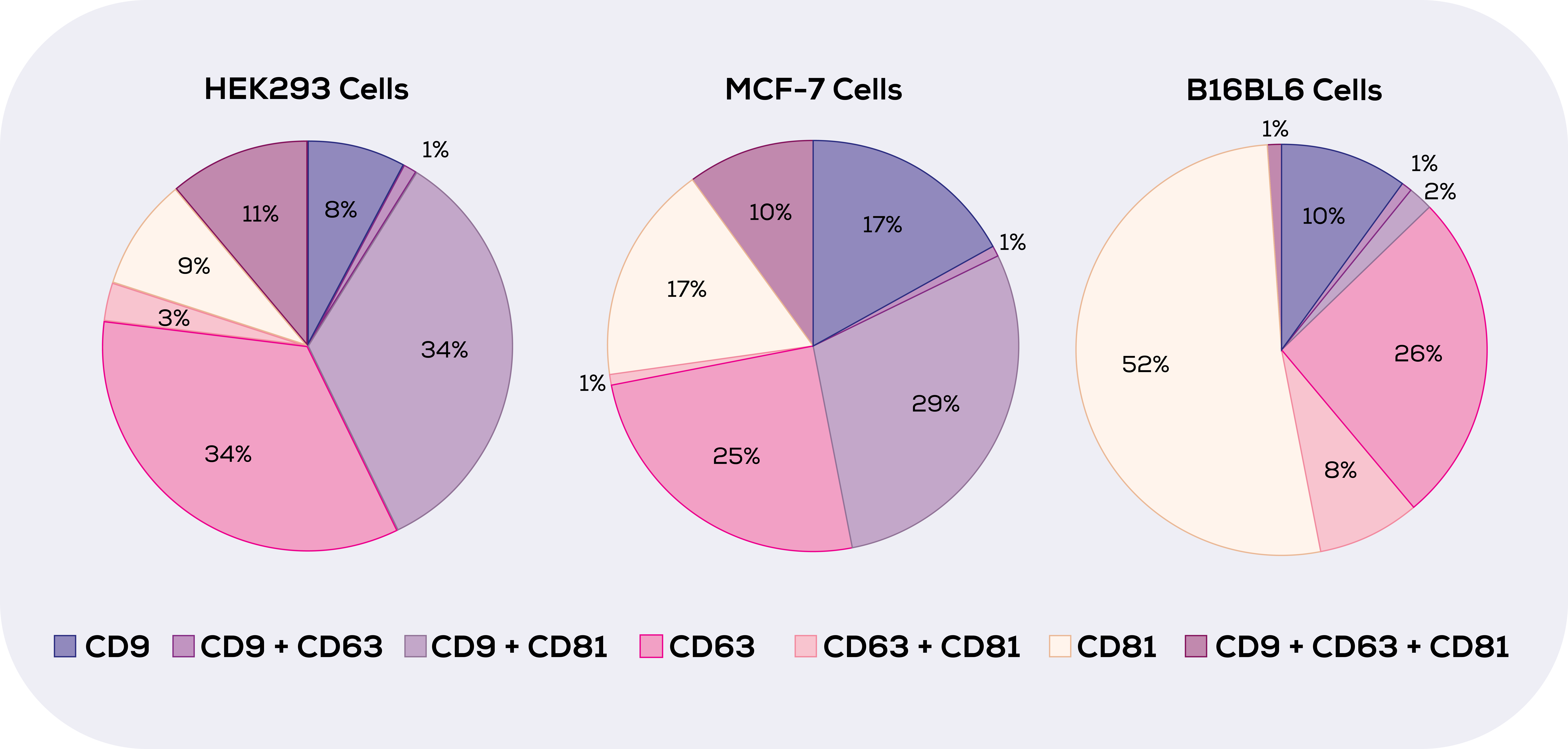
Other fluorescent labelling strategies include the use of proteins fused with green fluorescent protein (GFP) or similar fluorescent proteins (e.g., mCherry) through genetic modification of cell lines or organisms. This protein-associated labelling strategy – usually labelling of tetraspanins such as CD9, CD63 and CD81 – relies on perhaps inaccurate assumptions being made about the proteome of the EVs of interest. A study by Han et al (2021) found that none of these three tetraspanins were found on all EVs (or even close to all) in any of the three cell types investigated.5 As you can see in the visualisation of this in Figure 1, not only did no tetraspanin label all EVs, but the tetraspanin landscape of EVs from different cells is vastly different. This means that not only will no EV marker-based labels be able to label all EVs, but that they will likely bias labelling.
However, despite the lack of (known) bias in lipophilic labelling of EVs, there are other problems with this most favoured of methods.
The problem with lipophilic dyes and an unexpected plot twist
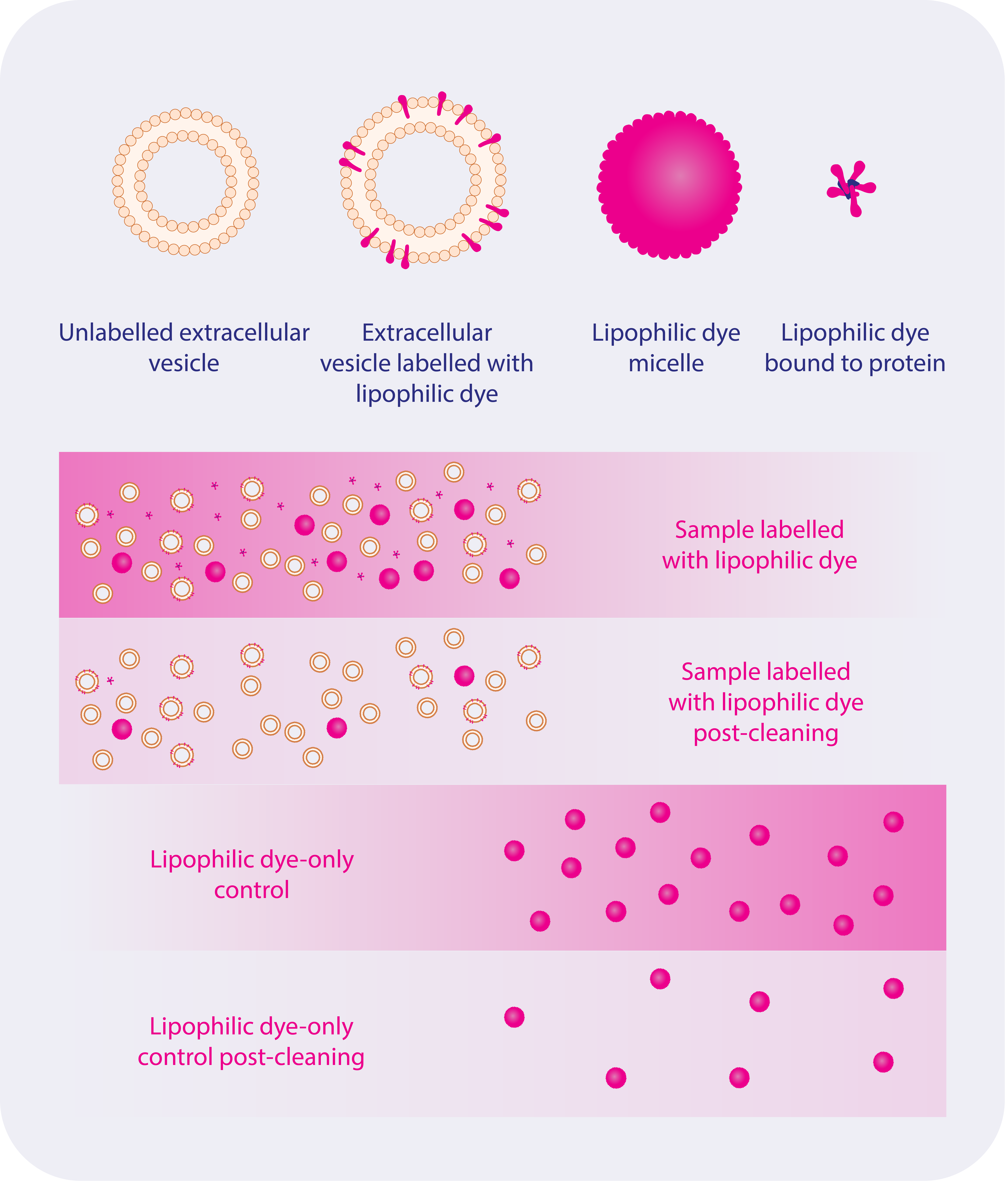
In 2018 a bombshell hit the EV world. Dominkus et al. published their study showing that PKH26 created non-EV particles which were internalised by cells.6 The problem? Many researchers at the time were using PKH26 to label their EVs. The study triggered them to subsequently question their results, sparking a flurry of studies on the topic. These studies have gone on to show that only a small proportion of dye intercalates into EVs, with the rest forming micelles, binding to protein or even forming large aggregates of unknown format (possibly also micelles), the first two of which are shown in Figure 2.6,7
This non-intercalated dye represents wasted potential signal, but it is autonomously (i.e., EV-independently) internalised into cells, masquerading as EV uptake.8 They are even internalised into punctate compartments in cells, likely the endosome-lysosome pathway.9 Again this masquerades as genuine EV uptake. As such, the most important step in EV labelling isn’t the labelling itself, but the removal of non-EV bound dye.
Removing non-EV intercalated lipophilic dye
Free dye has almost always been attempted to be removed in some form or other. I added protein in the form of BSA (that sticky protein of choice for biologists) to soak up as much dye as possible before using a desalting column to remove excess dye.10 However, many used ultracentrifugation, and still do. Using the ultracentrifugation method, the solution of EVs and excess dye is centrifuged at speeds conducive to pelleting EVs. However, this has been shown to potentially even contribute to the problem of micelles creation.6 Likely, my desalting columns only removed tiny contaminating free dye contaminants and left larger micelles and proteins present. To deal with the protein-associated dye, size exclusion chromatography (SEC) can be used, as this method has been shown to remove the majority of excess dye.9 Our SEC columns, qEV columns, have been successfully used to remove excess dye.11,12 Likely the successfulness of SEC to remove excess due will be (at least up to a point) positively correlated with the amount of protein (e.g., BSA) added to soak up excess dye, as SEC such as qEV columns are excellent at separating protein from EVs.
However, a small amount of excess dye does remain after SEC9, as it likely would with any single technique. This is because contaminating dye particles of different kinds may have the same size or density as EVs. This means that to remove all contaminating dye particles, combining a size based technique like qEV columns with a density based technique such as density gradient ultracentrifugation6 may be the be the best course of action. Whichever methodology you use, it is important to make a dye-only control which uses buffer in place of the EV solution, but otherwise is treated the same. In this way, you will be able to quantify and account for any non-specific dye uptake.
The future of EV labelling
Labelling EVs to study their uptake is not going to stop being important, instead it will likely increase in importance as more EV therapeutics move towards the market. In light of that, improving the efficiency and cleanness of labelling is essential. Our qEV columns are an integral part of this puzzle.
To find out more about qEV columns, visit the qEV overview page.










