The use of sensitive and reliable techniques for nanoparticle analysis lies at the heart of trustworthy data and scientific progress. When selecting an analytical technique for nanoparticle characterisation, considering the capabilities and limitations of each approach is an important step towards obtaining high-quality data.
Light scattering-based techniques such as dynamic light scattering (DLS) and nanoparticle tracking analysis (NTA) have been used widely across many fields of nanoparticle research, including extracellular vesicle research, environmental analysis, and nanomedicine. However, nanoparticle studies often require a far higher level of resolution than is provided by these traditional approaches. Tunable resistive pulse sensing (TRPS) has emerged as a high-resolution, single-particle technique, offering a way forward for more precise measurements of size, concentration, and zeta potential. If you’re unsure which technique is right for you, asking these three key questions can help point you in the right direction:
1) Can the analytical technique detect nanoparticle subpopulations of different sizes?
In some cases, your sample in question might be monodisperse – or very close to it. In other words, all particles might be uniform in size. In most real-world cases, however, nanoparticles of interest will be present in a range of dimensions. Nanoplastics, for instance, may fall within the size range of 1 nm to 1 µm, while extracellular vesicles (EVs) in a plasma sample may range from 30 nm to 150 nm. For the analysis of polydisperse samples, it is important to ensure your technique of choice can detect particles across a relevant size range.
To compare the ability of different techniques in nanoparticle size analysis, Vogel et al. (2021) applied multi-angle dynamic light scattering (MADLS), NTA, and TRPS to samples containing NIST‐traceable polystyrene particles (conjugated polymer nanoparticles, CPN) of four different sizes.
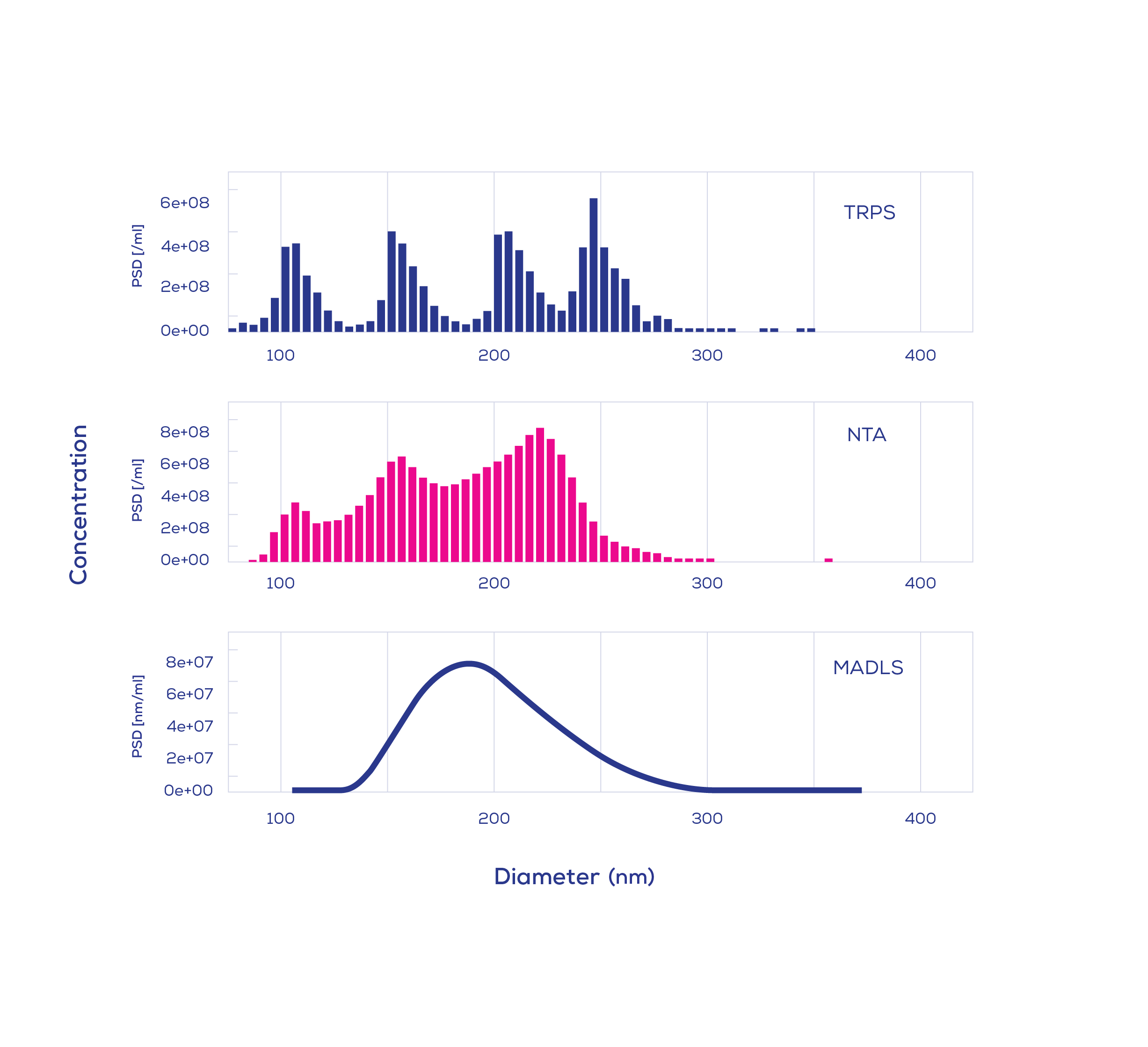
Despite the presence of four nanoparticle subpopulations, MADLS completely failed to resolve the quadrimodal mixture, while NTA only detected the presence of three subpopulations and failed to produce separate peaks. In contrast, all four subpopulations could be clearly identified using TRPS analysis. The same pattern was also shown by Caputo et al. (2021) whereby two different quadrimodal mixtures were assessed.
2) What is the smallest particle size that can be detected within a polydisperse sample?
When you’re working with polydisperse samples, it’s important to know the range of particle sizes that can be detected. In electron microscopy, for example, the presence of smaller particles can be underestimated – partly because they can be obscured by larger particles (Caputo et al. 2021). To assess the lower limits of particle size analysis, Vogel et al. (2021) analysed known samples of varying complexities.
When a trimodal mixture containing equal parts of CPN60, CPN100, and CPN150 (60, 100, and 150 nm, respectively) were applied to TRPS, NTA, and nano flow cytometry (nFCM), discrepancies between the different techniques were revealed (Vogel et al. 2021). While TRPS and nFCM fully resolved the subpopulations, NTA severely underestimated the proportion of 60 nm particles. With NTA, the proportion of 60 nm particles was estimated to be approximately 2%, rather than 33.3%.

3) How accurately can the technique measure total particle concentration?
Vogel et al. (2021) set out to answer this question by comparing techniques and their ability to measure the concentration of monodisperse samples; among them were MADLS, and single particle techniques NTA, TRPS and nFCM. TRPS and nFCM measurements were in line with nominal concentrations, while concentrations estimated by NTA was significantly higher than nominal concentrations. For MADLS, Vogel et al. (2021) observed an underestimation of total particle concentration. The overestimation of concentration by NTA has also been reported by others (Bachurski et al. 2019, Vestad et al. 2017).
Moving forward with precision and confidence
To ensure you are obtaining the most accurate and reliable data possible, it is important to consider the capabilities and limitations of each analytical approach. In most areas of nanoparticle research, samples of interest are polydisperse – yet the resolution of nanoparticle subpopulations cannot be achieved sufficiently using light scattering-based techniques.
TRPS offers a more precise way forward for nanoparticle research, providing a way to confidently characterize complex samples with single-particle resolution.
Get in touch now to discover how TRPS can support your nanoparticle research