Urine is an easily accessible, rich source of diagnostic information, making it second only to blood as the most utilised biofluid in the clinic.1 It is unsurprising then that interest in the composition of urine led to one of the first discoveries of what we would refer to as extracellular vesicles (EVs) in a 1986 study by Wiggins et al., long before EVs were described in the literature.2 These nano-sized, lipid-bilayer encapsulated vesicles contain a wide variety of different RNAs, proteins and small molecules from the cell that the EV originated from. It is this black box-like property – and their ability to shield potentially labile molecules from degradation – that makes EVs so attractive in the hunt for biomarkers. Their widespread presence in biofluids only increases their potential diagnostic utility. As with all biofluids, the unique composition of urine influences the special considerations that are required for EV isolation – a key aspect of developing urine-based EV biomarkers.
The complex composition of urine
Urine is a uniquely complex biofluid. Alongside cells from the donor (e.g., urinary tract epithelial cells, blood cells), urine also routinely contains microorganisms such as bacteria and even viruses. Interestingly, this property of urine has proven useful during the SARS-CoV-2 pandemic, where viral material has been routinely detected in wastewater, allowing for outbreak mapping even in low prevalence areas.3,4 In fact, enveloped viruses such as SARS-CoV-2, influenza and HIV are all encapsulated in host cell membranes and may share biogenesis pathways with EVs.5 This can make it difficult to distinguish between the two. Some viruses are suitably similar to EVs in size and density that it may be unavoidable to co-isolate viral particles in patients with viral infections.6 Therefore, unless infection is the pathology under investigation, exclusion of those patients/participants with viral or bacterial infection should be considered for discovery studies.
Typically, protein levels in urine are around 1000-fold lower than in plasma.7 However, this can change drastically in disease. As such, depending on your area of research, urine protein levels could vary widely. Even in healthy individuals, the concentration and composition of urine varies depending on a number of factors, including sex, age, hydration status, diet and time of collection.8,9 A position paper by the Urine Task Force for the International Society of Extracellular Vesicles recommended the use of dipsticks for common urine parameters (e.g., pH, blood contamination) to quickly determine whether a sample is suitable for analysis.1 Additional recommendations for the normalisation of urine concentration (e.g., to creatinine concentration) are also given in the position paper.1 The composition of urine impacts upon storage, pre-processing and EV isolation in a number of ways, meaning that there are special considerations for EV studies using this biofluid.
Considerations for collecting urine for EV studies
The first consideration for the isolation of urine EVs lies in the collection process itself. There are two major sources of variation introduced by the process of urine collection: collection schedule and void volume. Urine is generally considered to be most concentrated (and, perhaps, more likely to contain more EVs) when collected from the first urination of the day. The degree to which this impacts upon the characteristics of EVs, however, is of some debate.10,11 24 hour urine collections (e.g., a pooled sample of all urinations over a 24 hour period) can also be undertaken to maximise sample volume and minimise variations across the day. The second consideration is when in the void of urine to take samples (i.e., from the beginning or mid-stream). As no consensus has been reached on the best practice for urine collection in EV studies, it is important to be consistent within each study. The exception to this rule is for the isolation of prostate EVs from urine, where it is known that digital rectal examination produces increased prostate EVs in urine collected immediately following the exam.12
Pre-processing urine for EV studies: method considerations
As with all biofluids, pre-processing is essential for urine. This pre-processing looks different for different biofluids, but a common first step is the depletion of whole cells using centrifugation. For urine, this could also include the pelleting of bacteria. However, for urine there is another consideration – uromodulin. Uromodulin (also known as Tamm–Horsfall protein) is an abundant urinary glycoprotein which polymerises into filaments within which EVs can become entrapped.13 These polymers form particularly well at the low temperatures (i.e., around 4oC) at which urine is usually kept prior to processing. This refrigeration is particularly important for urine as this sample is often collected by patients themselves, and so cannot be immediately processed. Uromodulin is highly expressed in some cases of renal cancer, perhaps conferring increased survival, according to data in the NIH National Cancer Institute Genomic Data Commons14, suggesting that greater emphasis on uromodulin removal may be important in renal cancer patients.
It has been shown that centrifugation at 2,000 x g for 30 minutes can pellet the majority of uromodulin without significant loss of EVs.15 This method may be useful when mass spectrometry will be undertaken for EV protein analysis, where depletion of non-EV proteins is of particular importance. Another reported method instead aims to dissociate the uromodulin polymer using reducing conditions, though this was found to impact upon EV protein marker detection by Western blot, likely due to protein conformational changes.15 In this study, tunable resistive pulse sensing (TRPS) was subsequently used to measure EV concentration and size.15 When the goal is to determine accurate EV size and concentration, therefore, the use of reducing conditions for uromodulin dissociation may be more suitable than centrifugation, as it avoids the risk of EV loss.
Storing processed urine samples prior to EV isolation
Following pro-processing, urine must often be stored before EV isolation can occur, sometimes for long periods of time in biobanks. In the literature, storage at -80oC has been recommended.10 Though, EV concentration was found to drop and EV size was found to rise over one year of storage regardless of whether EVs were stored at -20oC, -80oC and -196oC, as measured by TRPS.16 This study found that nanoparticle tracking analysis was unable to ascertain size increase at any temperature but -20oC, highlighting TRPS as a more sensitive technique in this context.16
Isolating EVs from processed urine samples
Turning next to isolation of EVs, the method most commonly used for urine samples is ultracentrifugation, though this technique is highly variable and non-standardised in the field1, leading to the likely isolation of different subpopulations of EVs between studies. Urine also contains a considerable amount of albumin, meaning that good separation of EVs from the soluble protein fraction of processed urine is essential to prevent contamination of isolates. Recently, studies have begun to compare methodologies in an aim to maximise purity and reproducibility for EV isolation from urine.
One study found that size exclusion chromatography (SEC) using qEV columns achieved greater EV purity (i.e., lower protein contamination), than a precipitation-based method.17 Interestingly, another study found that ultracentrifugation was poor at isolating EVs from urine, resulting in the lowest concentration of EVs from 5 compared techniques.18 SEC using the qEVoriginal/70 nm columns isolated a high concentration of EVs in the same study, and showed the highest particle-to-protein ratio, indicating high purity.18 Both precipitation and ultracentrifugation resulted in low purity. qEV EV isolates were also free of uromodulin in this study, whereas ultracentrifugation showed high levels of uromodulin contamination.18 Finally, levels of the EV tetraspanin markers were highest in isolates from qEV columns.18 Another study agreed with these findings, showing that qEVsingle/70 nm columns were superior for urine EV isolation, particularly for RNA studies, when combined with pre-concentration using ultrafiltration.19 Additionally, another study agreed in finding SEC to be superior.20 As such, the literature suggests that urine EV isolation by qEV is a superior choice to other methods. This is particularly important for EV biomarker research where purity is essential.
Urine-derived EVs: a goldmine of biomarkers?
Urine is easy to collect and can even be collected by patients themselves, making urine a potential ideal source of EV biomarkers. It is of no surprise then that the majority of the research into urine EVs has focused on their biomarker applications, and it was from this research that one of the urine EV success stories arose.
The mRNA transcripts for PCA3 and the fusion gene TMPRSS2:ERG were first identified as being predictors of prostate cancer in the early 2000s, and were identified as being present in the urinary EVs of prostate cancer patients in 2009.21 Since then, this combination of mRNAs has been validated in numerous large studies, and now forms an FDA Breakthrough Device Designated diagnostic test with the ability to predict disease progression and guide the need for biopsy.22-24 Other biomarker studies have focused on diabetic nephropathy25,26, breast cancer27, cholangiocarcinoma28, pancreatic ductal adenocarcinoma29, nephritis associated with lupus30-32 and many more pathologies. It is of no surprise then that studies into urine EV biomarkers have been growing year on year, as shown in Figure 1. However, improvements to the standardisation and methodology of urine collection, pre-processing and EV isolation are likely important for more EV success stories from this rapidly increasing body of research – and answering the questions in the field may help with this process.
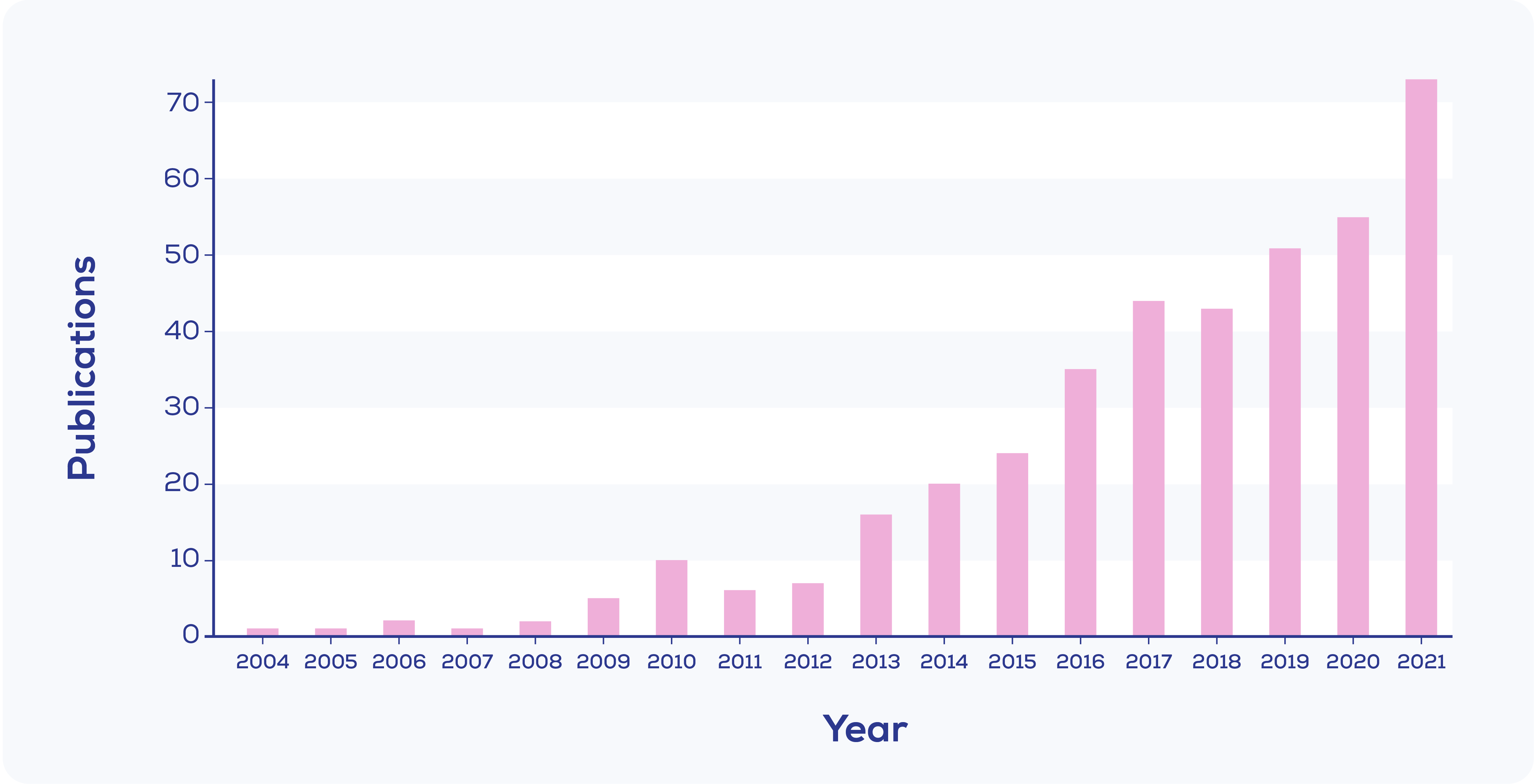
Figure 1. Publications mentioning 'urine' and either 'extracellular vesicles' or 'exosomes' in their title, with the word 'biomarker' in their title or abstract. Created from Web of Science data.
Key questions for urine EV research
Urine EV research is fairly advanced when it comes to the number of studies, yet some key questions remain unanswered.
- Which method is best for removal of uromodulin? Uromodulin removal is key to maximising EV yield and purity and preventing possible subpopulation depletion. Both centrifugation and reducing conditions show promise, but the optimal conditions for preservation of functional EVs in the sample have yet to be established.
- What is the best normalisation method of urine EV studies? Urine concentration and content varies widely with patient factors ranging from hydration to health status. A tested, agreed upon panel of normalisation factors would benefit reproducibility in the field.
- Will the field move away from ultracentrifugation and towards better technologies? Comparative research in the field of urine EVs has shown that isolation using SEC outperforms isolation using ultracentrifugation. Given the lack of concordance in ultracentrifugation protocols, moving to a standardised commercial workflow with the qEV isolation system would likely improve concordance between biomarker studies.
The identification of biomarkers from urine has been particularly successful to date, yet there is still a lack of urine EV based diagnostics in the clinic. Improving standardisation of urine collection and normalisation combined with improved, streamlined EV isolation could help these biomarker studies make the jump from bench to bedside.









