Nanoparticles (NPs) are at the forefront of advancements across therapeutics and biotechnology, and high-resolution characterisation is critical for such developments. In particular, physical characterisation is critical to process optimisation and the pursuit of obtaining the desired and biologically relevant properties.
In a paper published by Vogel et al. (2017), the authors describe improvements of the robustness and reproducibility of zeta potential measurements using Tunable Resistive Pulse Sensing (TRPS), a single-particle analytical technique designed for measuring nanoparticle size, concentration, and zeta potential.
The role of nanoparticles in nanomedicine
NPs, or particulate materials within the size range of 1 to 100 nm, have been integral to the progress of many fields, including applications in imaging, medicine, and environmental contexts.
It is well known that specific physicochemical characteristics are heavily linked to NP size, shape and material. Therefore, the precise control and verification of NP properties is of utmost importance when you are trying to obtain homogenous NPs with a particular feature. This is harnessed in applications using gold NPs, for example, where gold nanorods of different aspect ratios (i.e., sizes) in solution have different colours and absorption properties.
Compared to some traditional biotherapeutics, nano-scaled materials provide numerous advantages in nanomedicine, where there have been reports of higher efficacy, lower toxicity, and convenient administration routes to name a few. Therapeutics-compatible NP materials predominantly comprise the widely used synthetic lipid vesicles like liposomes or polymer-based conjugates, which can be further mixed or functionalised to improve desired biological interactions like binding to other molecules or cell surfaces. Thus, NP-based drug-delivery systems are recognised to have many advantages including the ability to deliver several drugs in a single vehicle, targeted delivery and controlled drug release reducing frequency of administration. However, in the journey towards clinical application and with the usual multicomponent and multifunctional modality of NP-based drug carriers, pre-clinical high resolution NP characterisation is commonly under-performed and a more complex task to execute.
Despite this, NPs have demonstrated great utility, such as in the method development of capture or purification systems. The use of magnetic NPs, for example, has significantly advanced purification assays thanks to the convenient separation of unwanted analytes using magnetic devices. Also, the immobilisation of molecules to the NP’s surface (e.g., aptamers, proteins, antibodies, lectins, DNA, and RNA) can be harnessed for therapeutic development, or for further differential binding/separation of other molecules.
The importance of nanoparticle physicochemical properties
Regardless of the intended application, the challenge remains: high-resolution NP characterisation tools are needed to determine whether NPs have been precisely engineered to achieve the intended goal – whether that is a unique biological nanointeraction, a modification for method optimisation, or meeting a relevant regulatory requirement relevant to particle stability.
Of the many physicochemical properties, key parameters such as size, surface charge and hydrophobicity take the spotlight as main drivers of the resulting and distinctive characteristic. Among these, particle effective charge or zeta potential is often measured for NPs in suspension meant for pharmaceutical purposes, as it is not only a way to validate protocol modifications but also to assess particle stability, i.e., repulsion or aggregation between NPs. The zeta potential is defined as the electrostatic potential at the plane of shear between the particle-associated and the stationary part of the ionic double layer and depends on factors such as surface charge density of the particle, dispersion medium (polar or non-polar), type of electrolyte, salt concentration, and pH.
Unlike some methodologies assessing zeta potential in NPs that are based on ensemble techniques, TRPS is a single-particle electrokinetic measurement technique that provides substantially more information about subpopulations within a sample and the samples’ true polydispersity. TRPS is a physical-based technology that relies on NPs translocating through a nanopore, creating a temporary decrease in current. The characteristics of these current blockades are used to determine particle concentration, size and zeta potential by measuring the number of blockades, blockade magnitudes and blockade durations, respectively.
New methodology helps TRPS deliver high-resolution single-particle measurements
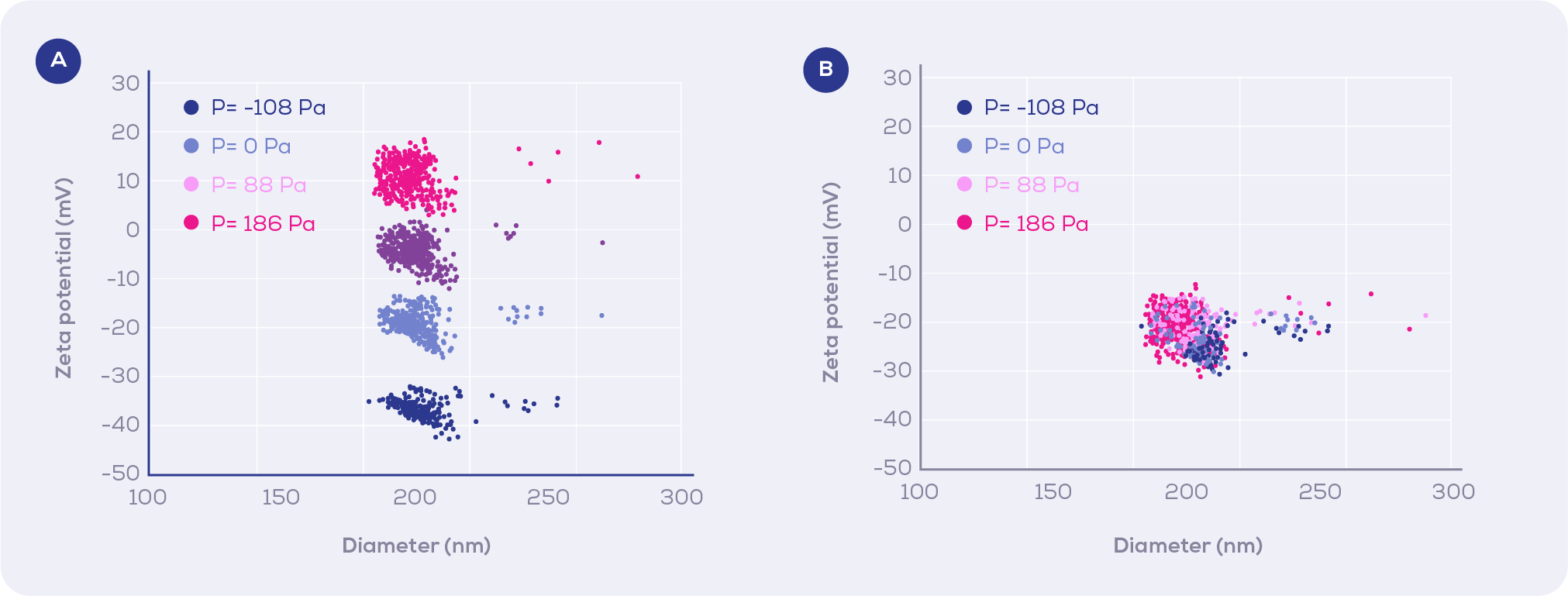
In this study, the authors improved the robustness and reproducibility of TRPS-based zeta potential measurements, by measuring the translocation duration of NPs as a function of voltage and pressure, instead of only voltage used by previous methods. This was achieved by including convection into the theoretical model by quantifying the convection from the slopes of the 1/T vs pressure linear curves used within the calibration method.
In this way, when convection is well defined, zeta potential measurements are independent of applied pressure (Figure 1b), as opposed to theoretical convection that results in varying zeta potential measurements when different pressures are applied (Figure 1a).
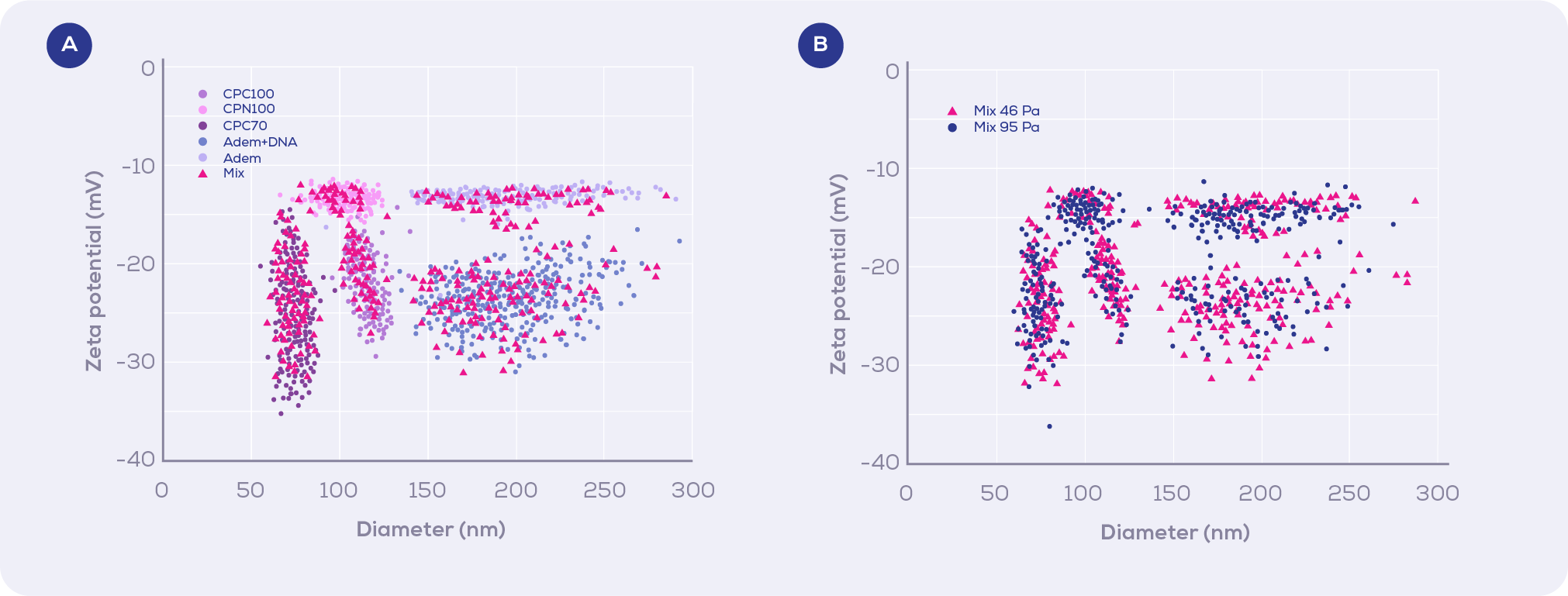
Moreover, the analysis of NP populations of multimodal zeta potential distributions demonstrated this approach can clearly discriminate and accurately resolve subpopulations with subtly different NP sizes and zeta potentials when run independently (Figure 2a) and as a mix (Figure 2b).
Accurate characterisation of liposomal formulations
Liposomes are a leading material class of commercially established and regulatory accepted drug-delivery carriers. Liposomal formulations usually contain mixtures of anionic and cationic liposomes to better mimic cellular membranes and improve their interaction, therefore, enable effective drug delivery. With the formulation having such a strong impact on the drug’s efficacy, zeta potential profiling becomes a critical step in tracking whether different lipids have been successfully mixed into liposomes.
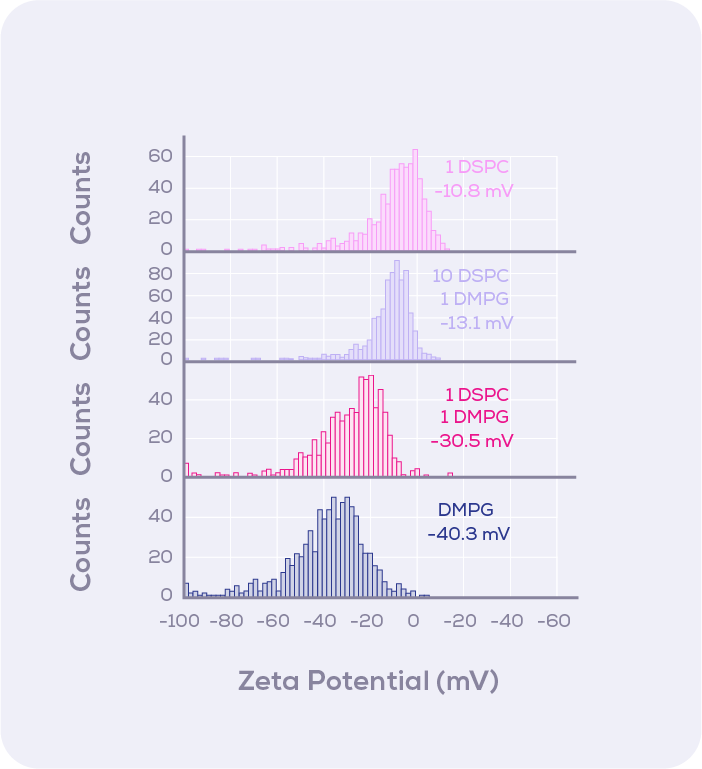
In this study, liposomal formulations containing different proportions of two types of lipids were profiled in their zeta potential distributions. Knowing the charge properties of each lipid, i.e., one zwitterionic (equal positive and negative charges) and the other negatively charged in this experiment, the mixing of liposomes was characterised through a high-resolution comparison of zeta potential (Figure 3).
Monitoring of oligonucleotide immobilisation on the nanoparticle surface
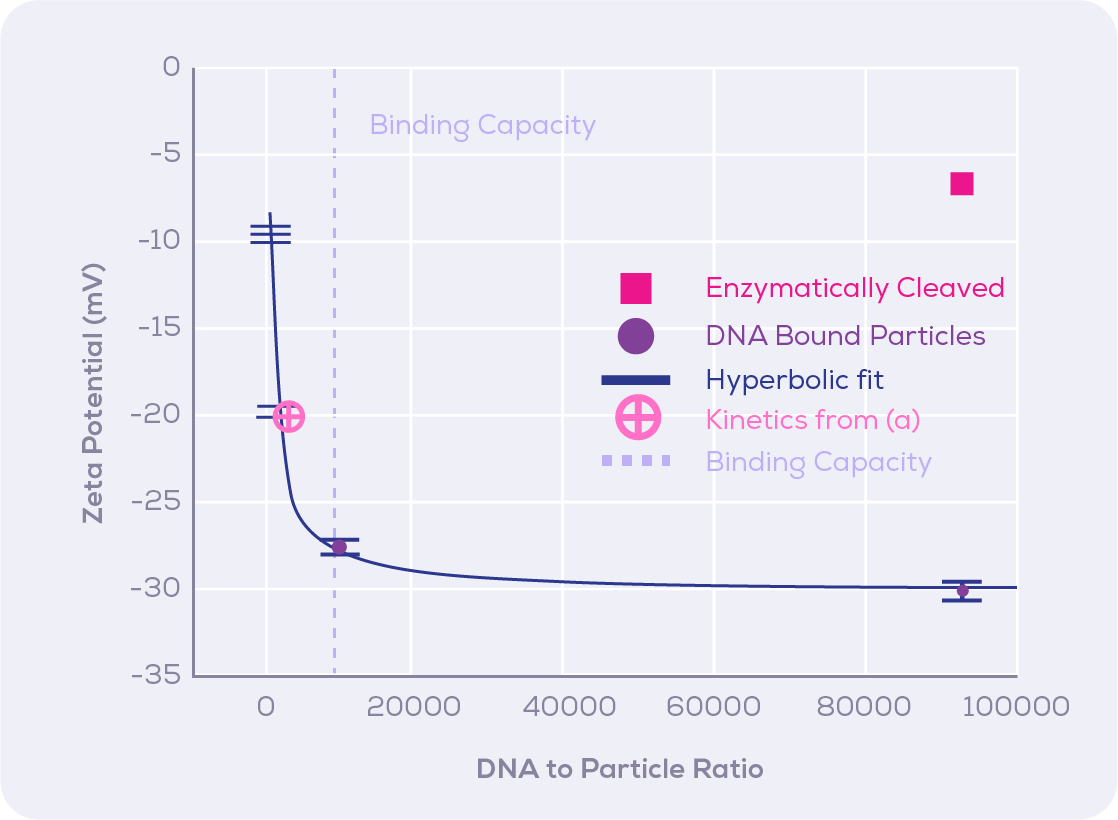
Immobilisation of oligonucleotides to surfaces has been utilised by numerous DNA-based bioassays: DNA sequencing, studies of DNA and protein interactions, biosensing and drug delivery. Attaching DNA specifically to NP surfaces further incorporates all the advantages previously described in bioassays. Monitoring the efficiency or timeline of DNA binding to NPs is critical for method optimisation and it requires a tool able to accurately characterise this dynamic. The improved method for TRPS-based zeta potential measurements was tested by monitoring the reaction kinetics of biotinylated DNA (35mer) onto streptavidin modified magnetic nanoparticles (MNP), while having MNPs with cleaved streptavidin (unable to bind to biotinylated DNA) as the control. The addition of DNA to MNPs led to a zeta potential change in MNPs that was quickly saturated at around 100 seconds. MNPs were then incubated with various DNA concentrations representing increasing DNA/MNPs ratios to show that the relationship of zeta potential and DNA/MNP ratio followed a hyperbolic behaviour (Figure 4). Differential calculations from the hyperbolic curve allowed the authors to determine the resolution or number of DNA molecules per MNP which can be distinguished via zeta potential measurements for different amounts of DNA loading. The authors mention that the resolution is specific to the DNA molecule used in the study and that the resolution could be improved by using DNA molecules with a higher charge.
In conclusion, modifications of zeta potential measurements presented here incorporate convection in a defined way and has electrophoresis dominating the particle motion, resulting in better overall resolution and accuracy in very dilute NP suspensions and a wider spectrum of zeta potentials. TRPS-based zeta potential measurements can be beneficial in numerous NP applications, granting high resolution single-particle analysis for this key and biologically relevant physicochemical property.






